Researchers in the laboratory of George Church, Robert Winthrop Professor of Genetics at Harvard Medical School and a core faculty member at the Wyss Institute for Biologically Inspired Engineering at Harvard, are working to produce the most modified bacterial genome to date. The researchers believe the method they developed will help others who are trying to make many edits at once to any organism’s genome.
Harvard Medicine News spoke with Church lab postdoctoral researcher Nili Ostrov and graduate student Matthieu Landon about how the research is being done and what the achievement will mean for the field of genome engineering.
HMS: What’s new about this work?
LANDON: Previously, our lab used precision genome editing technologies to replace about 300 instances of one particular codon each time it appeared throughout the genome of an E. coli bacterium. Codons are three-letter DNA sequences that code for amino acids, the building blocks of proteins. Replacing that single codon was a huge undertaking.
Here, we wanted to make over 60,000 genome changes so we could replace not just one but seven codons at once. Rather than gradually editing an existing genome, we did this by computationally designing and synthesizing an entirely new E. coli genome, with all the necessary genetic modifications already built in.
“We are pushing to unprecedented levels the technologies available for changing whole genomes. There is no other synthetic organism that has so many changes compared to its natural genome.” —Nili Ostrov
HMS: Did it work?
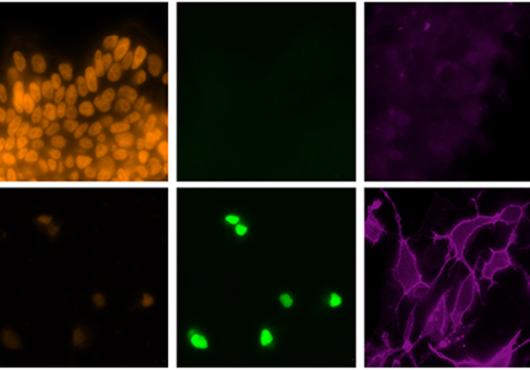
OSTROV: Though we are not done yet, so far it works very well. We constructed the synthetic genome in 87 overlapping segments and we are testing them one by one in living cells. Our goal is to confirm that, when we insert the recoded genes into a cell, the cell stays alive and expresses proteins normally. Then we will combine all the segments into a single fully-recoded genome.
LANDON: We’ve scrutinized about two-thirds of the genome so far (55 out of the 87 segments), and to our great happiness and surprise, we have found problems in only 13 of 2,229 genes tested. That is a very small number considering the extent of changes we made, which gives us confidence that our method is working and that we can complete the final organism.
HMS: Why not just test the whole genome at once?
OSTROV: If we had made all of the changes at the same time, and then the genome didn’t work, it would have been very difficult to pinpoint exactly which of the 4,000 altered genes was troublesome. On the other hand, testing individual genes would have taken a very long time.
LANDON: This is why we’ve chosen an intermediate segment size for testing. Working with these larger pieces of DNA allows us to test batches of genes at the same time—but not too many—so that we can quickly troubleshoot any problems. This was a critical aspect of our assembly method.

HMS: You’ve described this E. coli genome as “highly modified.” What does that mean?
OSTROV: One aspect is that we are changing the way the DNA code is being translated. Most organisms on Earth use 64 codons to translate their DNA into proteins. Our recoded E. coli is designed to use only 57 codons.
We are pushing to unprecedented levels the technologies available for changing whole genomes. There is no other synthetic organism that has so many changes compared to its natural genome.
HMS: Why would you want to do that?
LANDON: The main reason is that the bacterial strain we are trying to build will have new properties useful for medical and industrial applications that are very difficult or impossible to obtain through so-called punctual changes to the genome.
Examples of useful properties are virus resistance and genetic isolation. Many valuable compounds, including drugs such as insulin, are produced using bacterial strains. Usually, if a virus infects a batch of drug-producing bacteria, the whole plant has to be stopped and all the infected batches thrown away. Having virus-resistant bacteria could prevent such problems and potentially save millions of dollars.
OSTROV: You can also use synthetic organisms for basic science research. We don’t know of any other organism that uses only 57 codons, and it will be very interesting to investigate the effects of large-scale codon exchange.
HMS: How does removing codons help with something like virus resistance? It seems more intuitive that you would need to add DNA, not take it away.
OSTROV: That’s a good question. Removing codons is equivalent to removing genetic “letters,” changing the bacterium’s genetic language.
LANDON: Viruses hijack cells to replicate. To do this, they rely on the host cells having the same genetic code that they themselves have. By drastically changing the genetic code of our bacterial strain, we make it less compatible with viruses. We went down to 57 codons because to the best of our knowledge, most viruses that try to infect this strain will not be able to replicate.
HMS: What other possibilities does a reduced-codon organism provide?
LANDON: What you can do—and this is not something we address in this work, but is one of the long-term goals of synthetic biology—is repurpose the codons you have removed. A cell devoid of a codon will no longer know what it means, so we can now give the codon a new meaning—we can make it encode a new amino acid. These ‘nonstandard’ amino acids have now been used by many research groups to produce new types of proteins with diverse properties.
You can also use nonstandard amino acids to ensure that an organism is biocontained—that is, make it dependent on a substance that doesn’t occur in nature, so it’s unable to survive outside a controlled laboratory or industrial environment.
“It’s really the difference between editing a text—which we were doing quite efficiently before—and the ability to write a new text. Now we can write to the scale of an entire E. coli genome, which is very big.” —Matthieu Landon
HMS: You said there was another aspect to calling the genome highly modified?
OSTROV: Yes. It’s 100 percent synthetic. Most cells are daughters of parent cells, carrying a copy of the parent’s DNA. This organism’s DNA is going to be 100 percent chemically synthesized, not naturally replicated.
We designed the entire DNA code on the computer. We made that DNA by chemistry. Then we put it together in a living cell. That genome is a hybrid of our computational design and the E. coli genome blueprint.
LANDON: It’s really the difference between editing a text—which we were doing quite efficiently before—and the ability to write a new text. Now we can write to the scale of an entire E. coli genome, which is very big.
HMS: Some would say that’s a lot of power.
OSTROV: Like all other technologies, it needs to be properly controlled and it needs to be open to public scrutiny. We are very aware of the need to be responsible and do synthetic biology in a helpful, safe way. Biocontainment is a first step in this direction.
HMS: How does this work relate to discussions about “writing” a human genome?
OSTROV: It definitely shows we can write large genomes, but it also shows that it is not an easy thing to do. Even with an E. coli genome, which is a lot smaller and less complicated than a human’s, there are still many technological challenges we need to address. It’s not, ‘Oh, here. Synthetic organisms just work.’ These things are possible, but it’s a process.
HMS: What are you learning along the way?
LANDON: It’s hard to predict the biological effects of genome sequence changes. We have a lot of data that we have started to analyze—measures of the cell’s fitness, gene expression, and so on—and it will be very interesting to have the scientific community look at it.
Then, each problem we discover is usually something we haven’t anticipated during the design phase, which means it’s interesting on a biological level. We keep finding things that none of the current models explain.
OSTROV: What I find interesting is the way this project brings an engineering approach to biology. You want to engineer a cell, but biology is often unpredictable. The process of designing a whole genome, synthesizing and testing and then going back and fixing it, iteratively and a little more predictably, is very appealing to me. It’s a new way to do biology.
This interview was edited for length and clarity.
References:
Ostrov N, Landon M, Guell J, Kuznetsov G et al. Design, synthesis and testing toward a 57-codon genome. Science. Aug. 19, 2016. DOI 10.1126/science.aaf3639
Napolitano MG, Landon M, Gregg CJ, Lajoie MJ et al. Emergent rules for codon choice elucidated by editing rare arginine codons in Escherichia coli. PNAS. Sept. 20, 2016. DOI 10.1073/pnas.1605856113