Image: iStock/wildpixel
In pursuit of a novel tool for the research and treatment of celiac disease, Harvard Medical School scientists at the Mucosal Immunology and Biology Research Center (MIBRC) at Massachusetts General Hospital have validated the use of intestinal organoids.
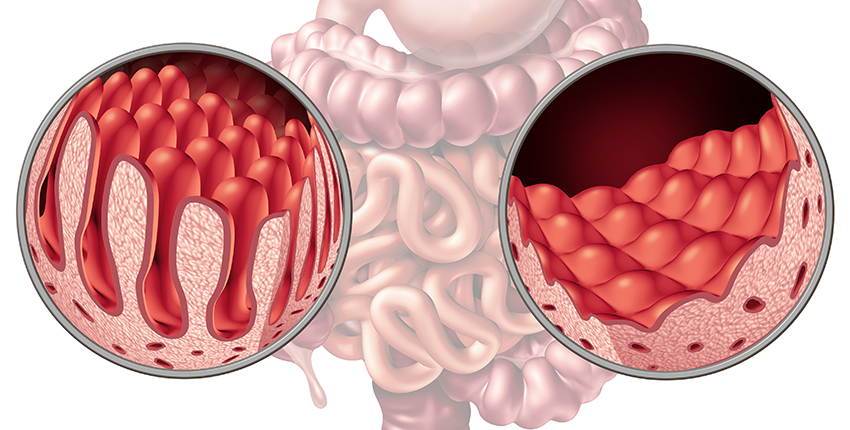
These three-dimensional tissue cultures are miniature, simplified versions of the intestine produced in vitro. Taking tissue from duodenal biopsies of celiac and non-celiac patients, researchers created the “mini-guts” to explore how the gut epithelium and microbiota-derived molecules respond to gluten, a complex class of proteins found in wheat and other grains.
“We currently have no animal model that can recapitulate the response to gluten that we see in humans,” said Stefania Senger, HMS instructor in pediatrics at Mass General and co-senior author of the study published on May 7 in in Scientific Reports.
“Using this human tissue model, we observed that intestinal organoids express the same molecular markers as actual epithelium in the celiac tissue, and the signature gene expression reflects the functional differences that occur when the epithelia of celiac disease patients are exposed to gliadin,” said Senger. Gliadin and glutenin proteins are main components of gluten.
Celiac disease is triggered when genetically predisposed individuals consume gluten. The condition affects approximately 1 percent of the U.S. population. Based on current data, the onset of celiac disease is thought to be preceded by the release of the protein zonulin, which is triggered by the activation of undigested gliadin to induce an autoimmune response. This leads to increased intestinal permeability and a disrupted barrier function. Novel evidence suggests that the microorganisms in the gastrointestinal tract may play a role in the onset of celiac disease.
Earlier studies from the MIBRC group and others have shown that human organoids “retain a gene expression that recapitulates the expression of the tissue of origin, including a diseased state,” the authors wrote. Through RNA sequencing, the new findings validate the organoid model as a “faithful in vitro model for celiac disease,” Senger said.
Using whole-transcriptome analysis, the researchers identified 472 genes regulated differently in organoids reflecting celiac disease than in non-celiac control organoids. These included novel genes associated with epithelial functions related to the pathogenesis of celiac disease—including gut barrier maintenance, stem cell regeneration and innate immune response. A second finding of the study shows that bioproducts derived from gut microorganisms can be employed to modify the epithelial response to gluten, a finding that could lead to future treatment strategies.
“These results confirm our hypothesis that genes and exposure to gluten are necessary but not sufficient, since changes in both the composition and function of the gut microbiome are also needed to switch from genetic predisposition to clinical outcome, as shown by our data,” said Alessio Fasano, HMS professor of pediatrics at Mass General, director of MIBRC and co-senior author of the paper.
“We believe our observations represent a major shift in the study of celiac disease,” Senger added. “We are confident that with adequate funding we could achieve major goals that include the development and implementation of high-throughput drug screenings to quickly identify new treatments for patients and expand the organoid repository to develop more complex models and pursue personalized treatment.”
Support for the study includes the National Institutes of Health (grants RO1 DK104344-01A1 and 1U19 AI082655-02) and the Egan Family Foundation.
Adapted from a Mass General news release.