Scientists have wondered whether somatic, or non-inherited, mutations play a role in aging and brain degeneration, but until recently there was no good technology to test this idea.
A study published online Dec. 7 in Science, led by researchers from Boston Children’s Hospital and Harvard Medical School, used whole-genome sequencing of individual neurons to find strong evidence that brain mutations accumulate as we age.
They also found that mutations accumulate at a higher rate in people with genetic premature aging disorders that cause early brain degeneration.
“It’s been an age-old question as to whether DNA mutations can accumulate in neurons—which usually don’t divide—and whether they are responsible for the loss of function that the brain undergoes as we get older,” said Christopher A. Walsh, the Bullard Professor of Pediatrics at HMS, chief of the Division of Genetics and Genomics at Boston Children’s and co-senior author of the paper.
“It hasn’t been possible to answer this question before, because we couldn’t sequence the genome of a single cell, and each mutation accumulated is unique to each cell,” he said.
Testing neurons one by one
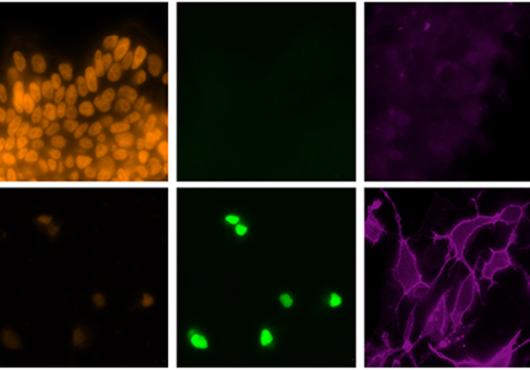
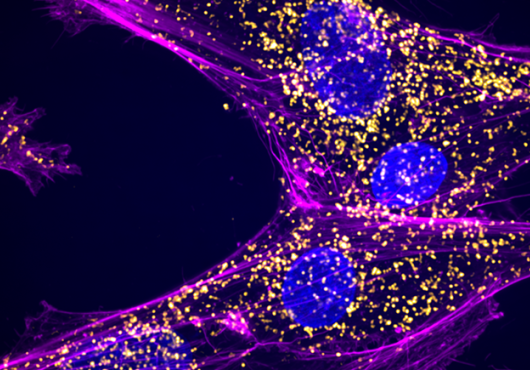
The research team tested DNA from 161 single neurons, taken from postmortem samples from the NeuroBioBank at the National Institutes of Health. The samples came from 15 neurologically normal people aged 4 months to 82 years and from nine people with one of two accelerated aging and early-onset neurodegenerative disorders, Cockayne syndrome and xeroderma pigmentosum.
Using the latest experimental and data analysis techniques, the team was able to detect mutations as small as single-letter changes in each neuron’s genetic code. Each cell had to have its genome amplified by generating a multitude of copies before its DNA sequence could be determined, and a large amount of data had to be analyzed.
“Because many experimental artifacts arise during the single-cell experiments, a new computational method that can distinguish true mutations from the experimental noise was critical to the success of the project,” said Peter Park, professor of biomedical informatics at HMS and the paper’s other co-senior author.
The neurons tested came from two areas of the brain implicated in age-related cognitive decline: the prefrontal cortex, the part of the brain most highly developed in humans, and the dentate gyrus of the hippocampus, a focal point in age-related degenerative conditions like Alzheimer’s.
In neurons from the neurologically normal people, the number of genetic mutations increased with age in both brain areas. However, mutations accumulated at a higher rate in the dentate gyrus. The researchers think this may be because the neurons have the ability to divide, unlike their counterparts in the prefrontal cortex.
In neurons from the people with Cockayne syndrome or xeroderma pigmentosum, there was an increase in mutations in the prefrontal cortex over time—more than twice the normal rate. Additionally, the researchers found that the portions of the genome that neurons used the most accumulated mutations at the highest rate, with help from collaborators at WuXi NextCODE.
The aging genome
The researchers coined the term “genosenium”—combining the concepts of the genome and senescence or senility—to capture the idea of gradual and inevitable accumulation of mutations contributing to brain aging.
The mutations themselves fell into three categories.
“We were able to take all the mutations we found and use mathematical techniques to deconstruct them into different types of DNA changes,” said Michael Lodato, one of six co-first authors on the paper. “It’s like hearing an orchestra and teasing out the different instruments.”
One category of clock-like mutations was strictly aging-related, accumulating like clockwork in both brain areas independent of disease status. Another type did not correlate with age except in the dentate gyrus, where mutation numbers in dividing neurons did increase over time.
The third type was associated with oxidative damage to DNA and faulty DNA repair; it increased with age and was seen in high numbers in Cockayne syndrome and xeroderma pigmentosum neurons, and to a lesser extent in normal neurons.
“This last finding convinced me I need more antioxidants,” quipped Walsh, who is also a Howard Hughes Medical Institute Investigator. “Overall, it raises a question as to whether neurodegenerative diseases are like cancer, relating ultimately to DNA mutation.”
The researchers are now turning their sights on other neurodegenerative disorders.
“The technology we used can be applied to any degenerative disease of the brain,” Walsh said.
Michael Lodato, Rachel Rodin and Michael Coulter of Boston Children’s and Craig Bohrson, Alison Barton and Minseok Kwon of HMS were all co-first authors on the study. Other coauthors were: Maxwell Sherman, Carl Vitzthum and Lovelace Luquette of HMS; Chandri Yandava, Pengwei Yang and Thomas Chittenden of the WuXi NextCODE Advanced Artificial Intelligence Research Laboratory; and Nicole Hatem, Steven Ryu and Mollie Woodworth of Boston Children’s.
The study was supported by the National Institutes of Health (K99 AG054749 01, F30 MH102909, 1S10RR028832-01, T32HG002295, U01MH106883, P50MH106933, R01 NS032457, U01 MH106883), the Harvard/MIT MD-PHD program, the Stuart H.Q. and Victoria Quan Fellowship in Neurobiology, the Allen Discovery Center program through The Paul G. Allen Frontiers Group and the Howard Hughes Medical Institute.
Adapted from a Boston Children’s news release.